EBVOSC : From raw biodiversity data to operational indicators through the Essential Biodiversity Variables
[Project accepted as a GOSC case study]
- Keywords
- Introduction
- Significance of the case study
- Research challenges and societal benefits
- Data requirement for the case study
- Statement of the problem(s) that need to be addressed by GOSC
- Engagement with the GOSC Initiative
- Deliverables
- Workshop SFE² GFO EEF
- Workflow examples
Keywords
Essential Biodiversity Variables, FAIR, workflow, ecoinformatics, metadata, Galaxy-E, GEO-BON, biodiversity observatories, Ecological Metadata Language
Introduction
Data integration in biodiversity science is complex, essentially because framework harmonizing data and methods is lacking. Getting interoperable data from raw, heterogeneous and scattered datasets to measure and understand the spatio-temporal dynamics of biodiversity from local to global scales is both necessary and challenging. Essential Biodiversity Variables (EBVs) represent a relevant framework for identifying appropriate data to be collated and for creating and implementing analytical workflows, from raw data to EBV data products.
Our aim is to operationalize EBV indicators by targeting the highest levels of FAIRness (Findable, Accessible, Interoperability, Reusable) for both data and source code implementation, so that data and tools can be widely shared and reused.
A number of open standards, tools, platforms used by international infrastructures. In particular, we are already engaged with the Galaxy platform initiative for source code management and use; the DataOne network of data catalogs; the Ecological Metadata Language standard for data management; the 2021-2023 BiodiFAIRse GO FAIR IN roadmap; and GEO BON’s roadmap (“Improve the acquisition, coordination and delivery of biodiversity observations and related services to users including decision makers and the scientific community”). In relation with the GOSC vision, EBVOSC will seek to utilize, contribute to and ensure interoperability with these initiatives.
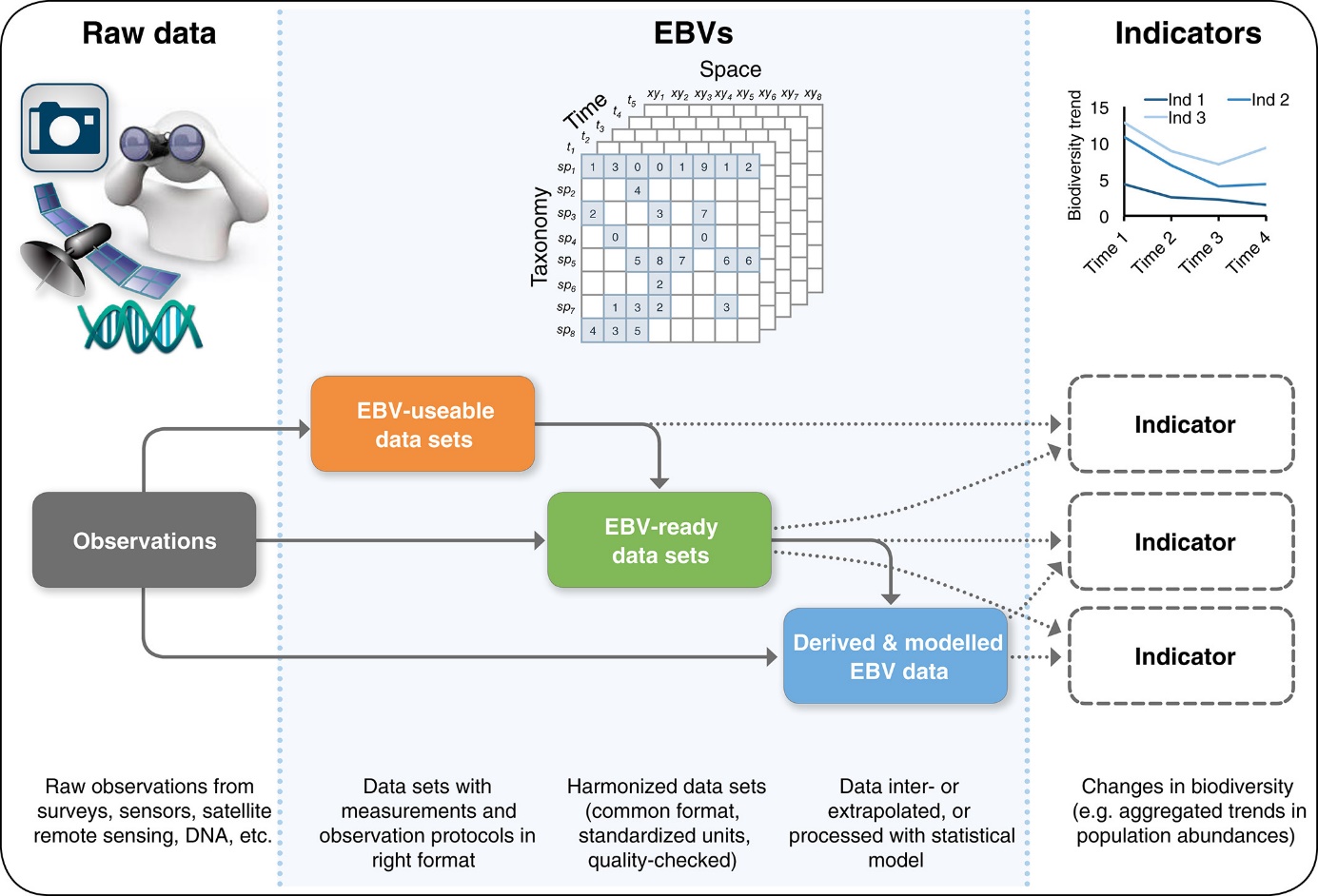
from Kissling et al, 2017

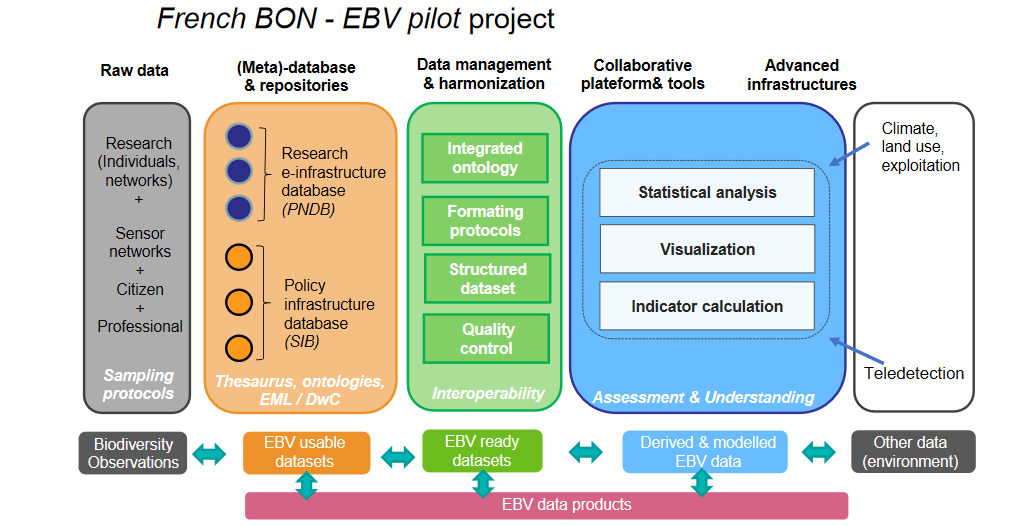
Significance of the case study
The EBVOSC case study aims to demonstrate that a better mobilization of such data can readily generate EBVs and associated biodiversity indicators through automated and regular updating.
For the biodiversity scientific community, EBV operationalization is a hot topic that raises several IT challenges (data structuration and sharing, source code review standardization and dissemination), thereby delaying our ability to respond quickly to face current biodiversity and climate emergencies. EBVOSC will provide an open and transparent comprehensive EBV operationalization pilot addressing these issues.
For the broader research community, EBVOSC will build on existing international standards, approaches and initiatives regarding data and workflows, thus benefiting communities in life, climate, and earth sciences as well as the humanities community, by linking biodiversity indicators to socio-economic measurements.
For society and stakeholders, operationalizing the EBV concept in a FAIR and transparent way is extremely important for people’s awareness on the biodiversity and climate crises through trusted indicators.
Research challenges and societal benefits
In line with international goals, measuring biodiversity state and dynamics in a transparent, reproducible and harmonized way, in line with driving forces and human pressures would have genuine societal benefits. By detecting change at species, population or community up to socio-ecosystems, they may allow appraising and reporting key and robust information at national and international levels (CBD, IPBES).
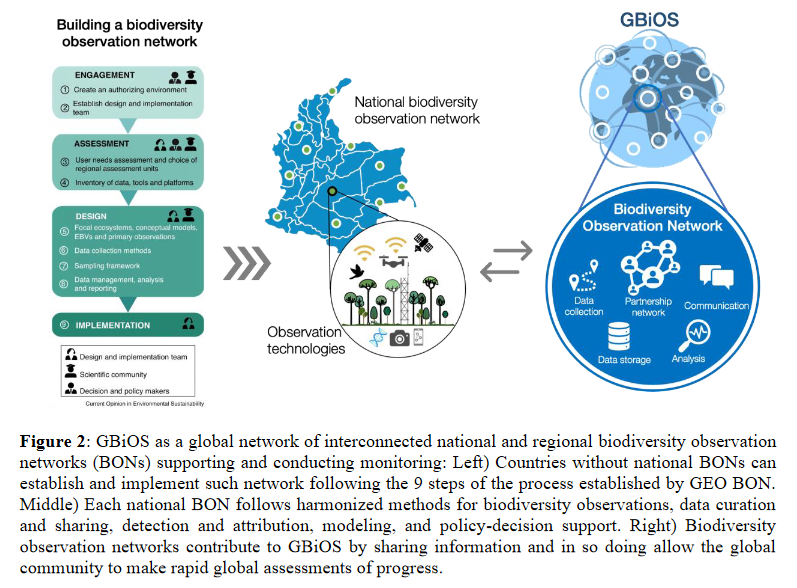
from Gonzalez et al., 2022 https://geobon.org/wp-content/uploads/2022/12/GBiOS_brief.pdf
Data requirement for the case study
Data from several biomes, both within and between BONs, will be gathered to demonstrate the portability and the reusability of EBV workflows. Extensive and well structured datasets (in particular nationwide surveys from each national BON) are candidate data for immediate EBV operationalization. Nevertheless, harmonization efforts would be required to make such data fully interoperable & reusable based on the highest degree of FAIRness.
Statement of the problem(s) that need to be addressed by GOSC
Dealing with multiple, scattered and heterogeneous data collection systems at all scales from gene to ecosystems, measuring biodiversity over large scales remains particularly complex. Based on existing approaches used by other domains such as climate or earth sciences, EBVOSC will propose methods to collate, harmonize and process contrasted in-situ data (e.g. field work and captor networks), and potentially together with remote-sensing data, by means of high performance computing tools and services. Existing efforts are not sufficient today to rapidly cope with the need of raw biodiversity data sharing and indicators production.
EBVOSC proposes an innovative way to address important challenges related to biodiversity indicators for facilitating existing biodiversity indicators production, and broad understanding.
Engagement with the GOSC Initiative
EBVOSC proposes to contribute substantially to GOSC working groups (Strategy, governance and sustainability / Policy and legal / Technical infrastructure / Data interoperability) from the biodiversity domain point of view. Nevertheless, EVBOSC is focusing on approaches and technologies that will also benefit other scientific domains.
Deliverables
- Highly reusable, transparent and accessible computational EBV workflows to be shared within and between BONs through “stacks” conda/container/Galaxy
- EBV data templates structured through EML metadata standard and terminological resources
- An evaluation process for the GOSC with stakeholders’ feedback
- Lessons and good practices for other OSCs.
Workshop SFE² GFO EEF

This projet has been presented at the joint meeting, Internatinal Conference on ecological sciences the 23th of November 2022, at Metz (France)
- un workshop : From raw biodiversity data to operational indicators through Essential Biodiversity Variables (cf. lien du congrès)
- Abstract : Because data integration with different ecological scales in biodiversity science is complex, the biodiversity community (scientists, policy makers, managers, citizen, NGOs) needs to build, a framework of harmonized and interoperable data from raw, heterogeneous and scattered datasets, in order to observe, measure and understand the spatio-temporal dynamic of biodiversity from local to global scale. One of the most relevant approaches to reach that aim is the concept of Essential Biodiversity Variables (EBV). Because we can potentially extract a lot of information from raw datasets sampled at different ecological scales, the EBV concept represent a useful leverage for identifying appropriate data to be collated as well as associated analytical workflow for processing these data. Thanks to FAIR data and source code implementation (Findable, Accessible, Interopability, Reusable), it is possible to make transparent assessment of biodiversity by generating operational indicators through the EBV framework, and help designing or improving biodiversity monitoring at various scales.
Based on the EBV, the French biodiversity data hub (“Pôle National de Données de Biodiversité” - PNDB ), is an e-infrastructure for and by researchers developing an integrated framework for 1) extracting EBV information from raw data using Ecological Metadata Language , (EML), 2) running reproducible ecological analysis through open-access workflows, and 3) producing biodiversity indicators for research, expertise and policy makers thanks to the Galaxy-Ecology collaborative platform .
In line with both the GO FAIR initiative and the GEO BON network , the PNDB is proposing a case study to focus on i) advancing conceptual developments related to EBV, such as the complementarities between EBVs and Pressure-State-Response frameworks (e.g. DPsIR) or the improvement of the research/expertise interface, ii) implementing EBVs for and with various communities (scientific research, expertise and policy makers), and iii) operationalizing EBV based on existing technologies (EML, Galaxy-Ecology). All of this will benefit various communities of biodiversity scientists.
-
- Authors : Coline Royaux (Sorbonne Université & MNHN), Jean-Baptiste Mihoub (Sorbonne Université), Olivier Norvez (FRB & MNHN), Sandrine Pavoine (MNHN), Dominique Pelletier (Ifremer), Aurélie Delavaud (FRB) & Yvan Le Bras (MNHN)
[The workshop presentation is available here]